Getting Started¶
You’re reading this because the API wasn’t cracking it and you would like to extend the framework for your own task or use some of the deeper layers of our codebase. This set of pages will take you from setting up the code for local development all the way to adding a new task or a new dataset to the framework. For using Torch Points3D as a library please refer to this section.
Installation¶
Install Python 3.6 or higher¶
Start by installing Python > 3.6. You can use pyenv by doing the following:
curl -L https://github.com/pyenv/pyenv-installer/raw/master/bin/pyenv-installer | bash
Add these three lines to your .bashrc
export PATH="$HOME/.pyenv/bin:$PATH"
eval "$(pyenv init -)"
eval "$(pyenv virtualenv-init -)"
Finaly you can install python 3.6.10 by running the following command
pyenv install 3.6.10
Install dependencies using poetry¶
Start by installing poetry:
pip install poetry
You can clone the repository and install all the required dependencies as follow:
git clone https://github.com/nicolas-chaulet/torch-points3d.git
cd torch-points3d
pyenv local 3.6.10
poetry install --no-root
You can check that the install has been successful by running
poetry shell
python -m unittest -v
Minkowski engine support¶
The repository is supporting Minkowski Engine which requires openblas-dev and nvcc if you have a CUDA device on your machine. First install openblas
sudo apt install libopenblas-dev
then make sure that nvcc is in your path:
nvcc -V
If it’s not then locate it (locate nvcc) and add its location to your PATH variable. On my machine:
export PATH="/usr/local/cuda-10.2/bin:$PATH"
You are now in a position to install MinkowskiEngine with GPU support:
poetry install -E MinkowskiEngine --no-root
Installation within a virtual environment¶
We try to maintain a requirements.txt file for those who want to use plain old pip. Start by cloning the repo:
git clone https://github.com/nicolas-chaulet/torch-points3d.git
cd torch-points3d
We still recommend that you first create a virtual environment and activate it before installing the dependencies:
python3 -m virtualenv pcb
source pcb/bin/activate
Install all dependencies:
pip install -r requirements.txt
You should now be able to run the tests successfully:
python -m unittest -v
Train!¶
You should now be in a position to train your first model. Here is how is goes to train pointnet++ on part segmentation task for dataset shapenet, simply run the following:
python train.py \
task=segmentation model_type=pointnet2 model_name=pointnet2_charlesssg dataset=shapenet-fixed
And you should see something like that
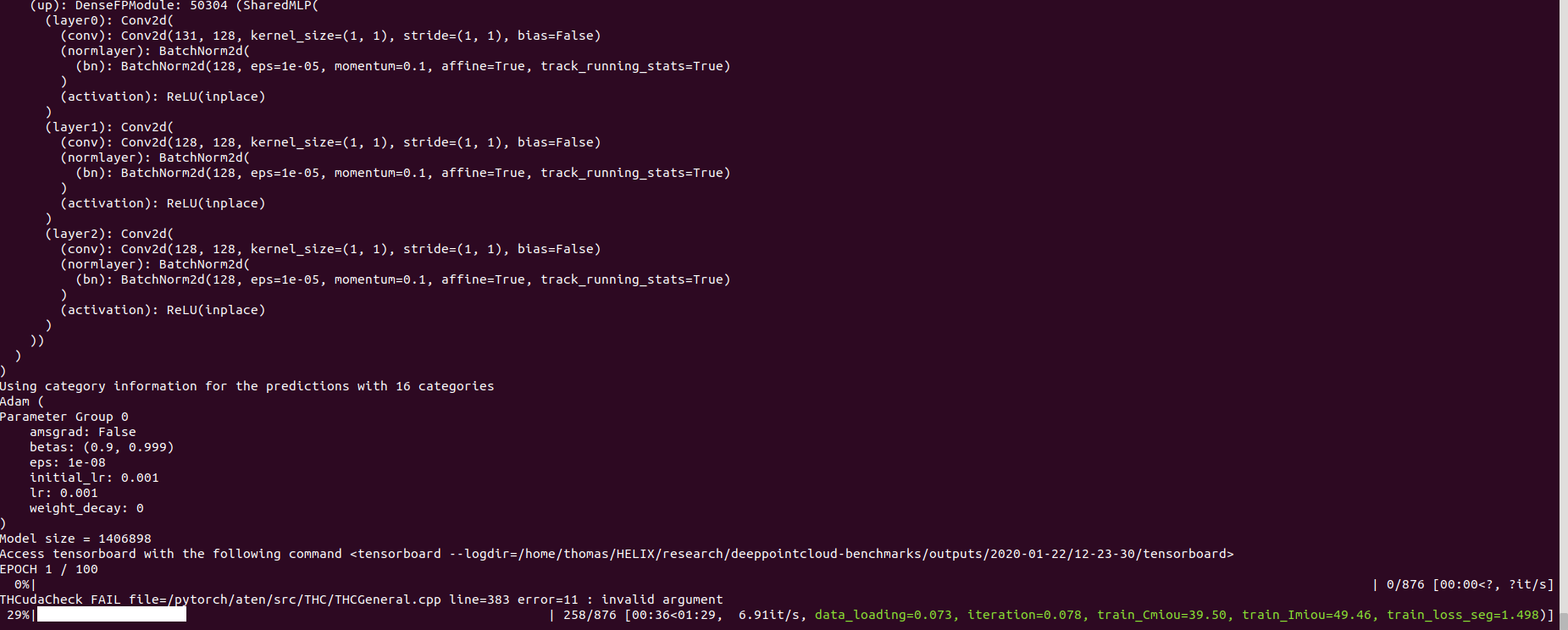
The config for pointnet++ is a good example starting point to understand how models are defined:
[512 + 64 + 32, 256, 256],
[256 + FEAT, 128, 128],
]
skip: True
mlp_cls:
nn: [128, 128]
dropout: 0.5
pointnet2_charlesmsg:
class: pointnet2.PointNet2_D
conv_type: "DENSE"
use_category: ${data.use_category}
down_conv:
module_name: PointNetMSGDown
npoint: [512, 128]
radii: [[0.1, 0.2, 0.4], [0.4, 0.8]]
nsamples: [[32, 64, 128], [64, 128]]
down_conv_nn:
[
[
[FEAT+3, 32, 32, 64],
[FEAT+3, 64, 64, 128],
[FEAT+3, 64, 96, 128],
],
[
[64 + 128 + 128+3, 128, 128, 256],
[64 + 128 + 128+3, 128, 196, 256],
],
]
innermost:
module_name: GlobalDenseBaseModule
nn: [256 * 2 + 3, 256, 512, 1024]
up_conv:
module_name: DenseFPModule
up_conv_nn:
[
Once the training is complete, you can access the model checkpoint as well as any visualisation and graphs that you may have generated in the outputs/<date>/<time> folder where date and time correspond to the time where you launched the training.
Visualise your results¶
We provide a notebook based on pyvista and panel that allows you to explore your past experiments visually. When using jupyter lab you will have to install an extension:
jupyter labextension install @pyviz/jupyterlab_pyviz
Once this is done you can launch jupyter lab from the root directory and run through the notebook. You should see a dashboard starting that looks like the following:
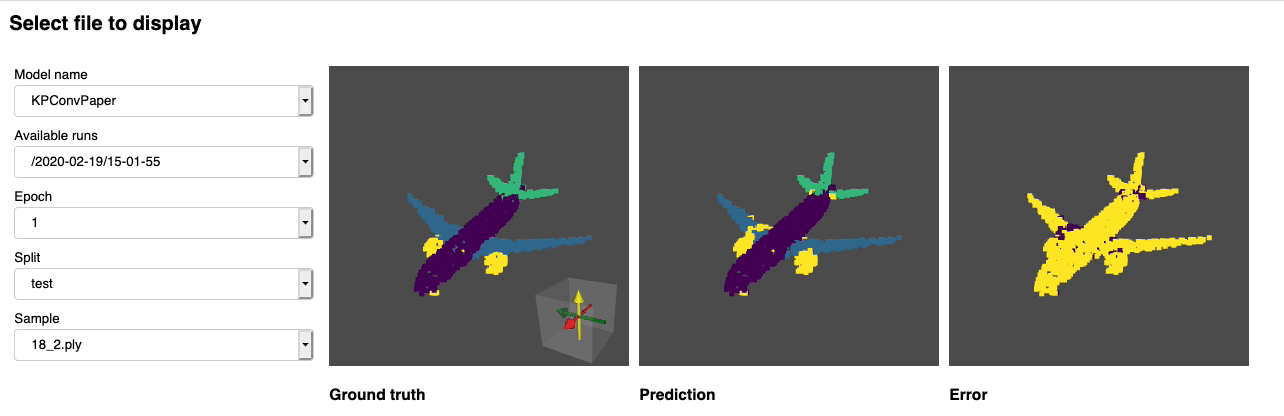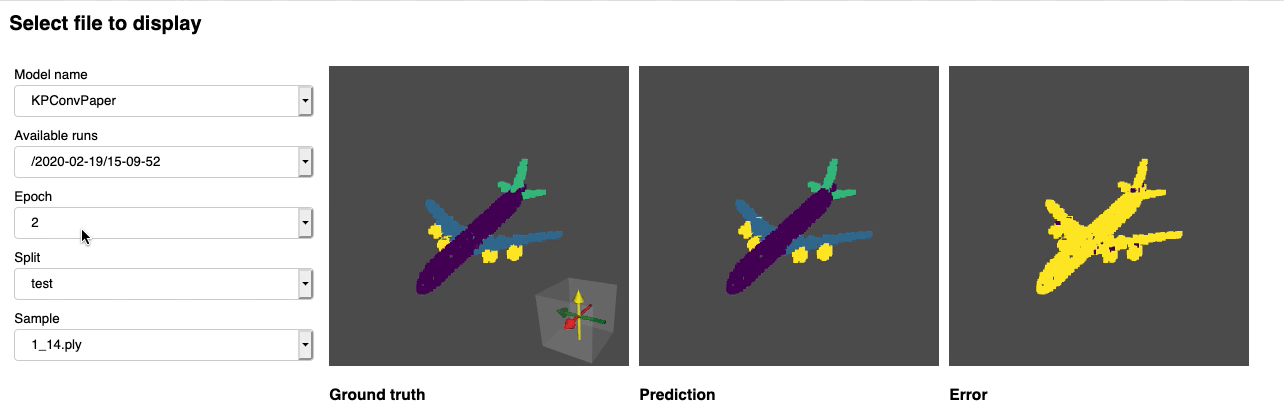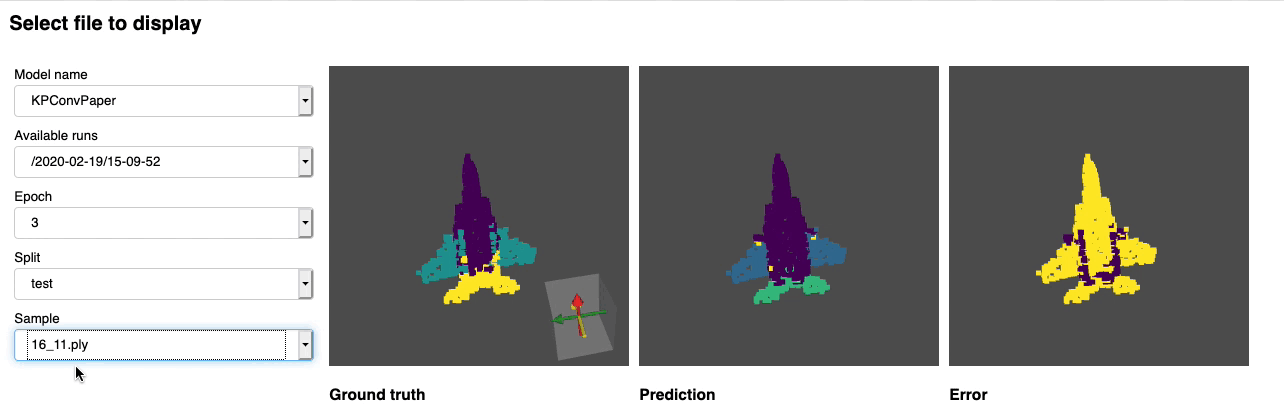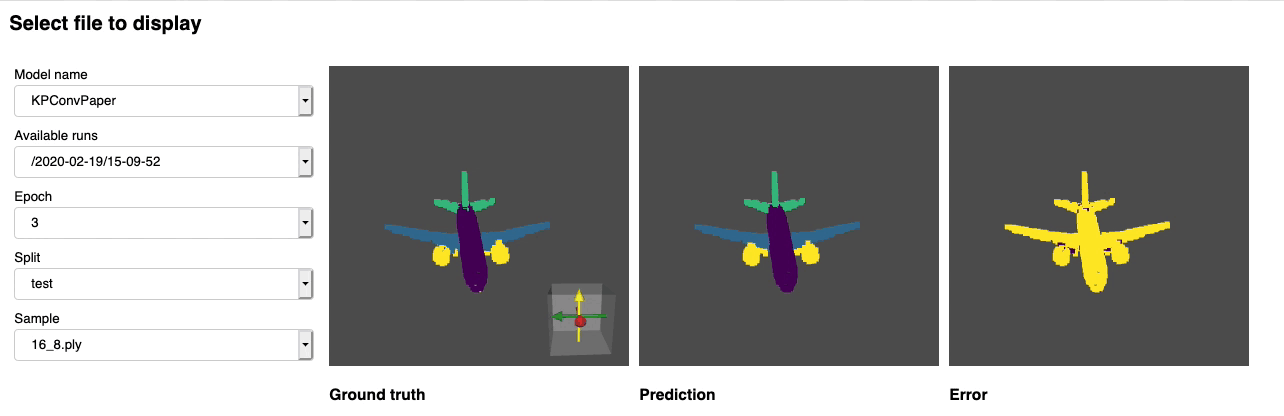
Project structure¶
The ambition of the project is to be a base for all point cloud related deep learning research. As such we wanted to make it scalable and also ensure that components could be reused. Below is the overall structure of the project:
├── benchmark # Output from various benchmark runs
├── conf # All configurations for training nad evaluation leave there
├── notebooks # A collection of notebooks that allow result exploration and network debugging
├── docker # Docker image that can be used for inference or training
├── docs # All the doc
├── eval.py # Eval script
├── find_neighbour_dist.py # Script that helps find the optimal number of neighbours for neighbour search operations
├── forward_scripts # Script that runs a forward pass on possibly non annotated data
├── outputs # All outputs from your runs sorted by date
├── scripts # Some scripts to help manage the project
├── torch_points3d
│ ├── core # Core components
│ ├── datasets # All code related to datasets
│ ├── metrics # All metrics and trackers
│ ├── models # All models
│ ├── modules # Basic modules that can be used in a modular way
│ ├── utils # Various utils
│ └── visualization # Visualization
├── test
└── train.py # Main script to launch a training
Note
As a general philosophy we have split datasets and models by task. For example, datasets has three subfolders:
segmentation
classification
registration
where each folder contains the dataset related to each task.